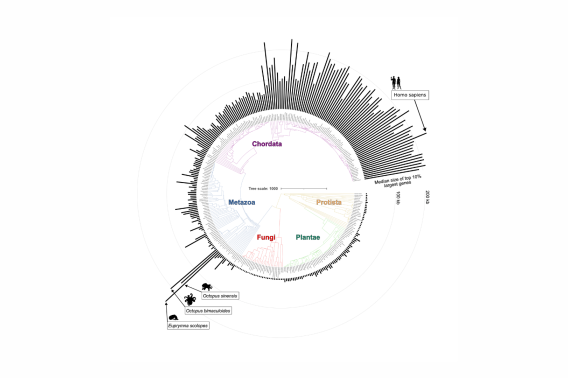
The role of gene complexity in the evolution and function of nervous systems
Many of the largest, most complex genes in the genome are enriched in the brain and are frequently mutated or misregulated in neurological diseases and disorders such as Alzheimer's disease, autism spectrum disorders, and Rett syndrome. This proposal outlines a detailed study of this distinct class of genes whose extreme sizes shape expression patterns, require unique regulatory mechanisms, and give rise to diversity as well as disease via shared processes. By comparing and contrasting the size, complexity, and regulation of neuronal genes in animals with simple nervous systems, such as the nematode, Caenorhabditis elegans, to animals with complex nervous systems, such as the cephalopod squid, Euprymna berryi, this study will characterize the functional consequences of gene size and complexity. The powerful genetic tools available in C. elegans will be leveraged to perform gene editing and a forward genetic screen to identify cell types capable of expressing large, complex genes, and to identify regulators of these genes. Newly available techniques for gene editing and aquaculture in the emerging model system, E. berryi, will be used to characterize long genes and their regulators in these deeply diverged invertebrates. Finally, single-cell transcriptomes of neuronal and non-neuronal tissues of both C. elegans and E. berryi will be sequenced with methods sensitive to isoform usage to define the relationship between gene size and molecular diversity among neurons. Together, the proposed research will illuminate the role of gene size and complexity in cell- and tissue-specific expression, and will identify novel regulators of large, complex neuronal genes in vivo. In doing so, we will gain a deeper understanding of how physical properties of the genome direct normal brain development and function, yet also contribute to neurological disorders and disease.